Inhaltsverzeichnis
Motion Dataminer
The datamining tool determines and extracts specific defined parameters (single values) from timeseries data, e.g. maximum, minimum or mean values of specific time-intervals (phases) or at specific time-positions (events).
XML Configuration file
An xml-configuration file defines, which values, from which timeseries, from where in the time space, should be extracted. Such configuration files are save in the projects subfolder „Datamining“ and have the suffix „.dml“.
Creation of a new configuration file
New configuration files can be created by the action „Create datamining filter“ in the Tools menue. A button to execute this action is typically available in the main toolbar. It opens an dialog which asks you for a name used as an filename for the created configuration file and an example trial.
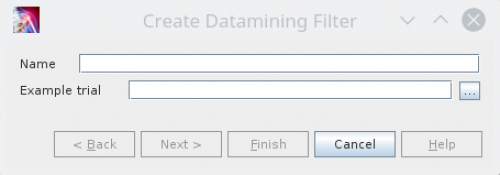 Hint: The example trial must be a file recognized as a trial. So a processed „.d3d“-file does not work.
Hint: The example trial must be a file recognized as a trial. So a processed „.d3d“-file does not work.
The action automatically creates the „expectedTrialContent“ based on the example trials content. In most cases this includes more than you want, so you have to delete the not needed parts.
Content of an configuration file
Have a look at the following example:
<datamining> <expectedTrialContent> <general defaultFileSuffix="ux"></general> <phaseTypes names="RCycle LCycle RCycleBW RCycleFW LCycleBW LCycleFW"></phaseTypes> <eventTypes names="RTO LG LTO RG RHS LHS GHS GTO"></eventTypes> <timeseries> <timeSerie components="1" name="LGlenohumeralEleDepDirCosAngle"></timeSerie> <timeSerie components="1" name="RShoulderAnteRetroHDCalDirCosAngle"></timeSerie> <timeSerie components="1" name="LShoulderIroAroAngleGlobe"></timeSerie> <timeSerie components="1" name="LShoulderIroAroHDCalAngle"></timeSerie> ... </timeseries> <properties></properties> </expectedTrialContent> <parameters> <parameter component="0" id="RGlenohumeralEleDepHDCalDirCosMax" name="RGlenohumeralEleDepHDCalDirCosAngle" type="max"></parameter> <parameter component="0" id="RShoulderEleDepHDCalDirCosMax" name="RShoulderEleDepHDCalDirCosAngle" type="max"></parameter> <parameter component="0" id="RShoulderAbdAddHdCalDirCosMin" name="RShoulderAbdAddHDCalDirCosAngle" type="min"></parameter> <parameter component="0" id="RShoulderEleDepHDCalDirCosMin" name="RShoulderEleDepHDCalDirCosAngle" type="min"></parameter> ... </parameters> <scopes> <scope id="0" name="RCycle" type="phase"> <variable id="RGlenohumeralEleDepHDCalDirCosMax" name="RGlenohumeralEleDepHDCalDirCosMax"></variable> <variable id="RTrunkObliquityHDCalMax" name="RTrunkObliquityHDCalMax"></variable> <variable id="RShoulderAbdAddHDCalProjMin" name="RShoulderAbdAddHDCalProjMin"></variable> <variable id="RShoulderAbdAddHdCalDirCosMin" name="RShoulderAbdAddHdCalDirCosMin"></variable> ... </scope> ... </scopes> <groups> <group fileSuffix="c3d" id="ShoulderAbdAdd" selected="true"></group> <group fileSuffix="c3d" id="ShoulderAnteRetro" selected="false"></group> <group fileSuffix="c3d" id="AroIro-0" selected="false"></group> <group fileSuffix="c3d" id="Schuerzengriff" selected="false"></group> <group fileSuffix="c3d" id="Kaemmen" selected="false"></group> ... </groups> </datamining>
Expected trial content definition
All possible data types which are expected in the trials and possibly should be extracted are listed as children of the <expectedTrialContent>-element.
The element <phaseTypes> defines a blank separated list of phase type names in its „name“-attribute. The element <eventTypes> equivalent for event-types.
The <timeserie>-element includes a list of <timeserie>-elements where the attribute „components“ defines the count of components of the timeserie and the attribute „name“ the name of the timeserie.
The element <properties> defines a list of properties as children. Each property defines a data type e.g. „String“ and a name:
<properties> <propertygroup name="TRIAL_INFO"> <property type="String" name="cycle_nr"/> <property type="String" name="pose_x"/> <property type="String" name="pose_y"/> <property type="String" name="pose_z"/> </propertygroup> </properties>
Parameter definition
After the <expectedTrialContent>-element you can find a list of so called parameters. Each parameter has an id, which is referenced in the scopes. The name-attribute specifies the name of a timeserie, as it is saved in the trials. The component attribute specifies the component. If the timeserie is a timeserie of only doubles the component is zero, in other cases it counts the component, starting by zero.
The most interesting is the „type“ attribute. Have a look at the following table to see, which types are currently available. Additionally types can be implemented easy, if needed.
| Type name | Description | Available for |
|---|---|---|
| max | maximum in the case of application to a „phase“, „value at“ in the case of an event | phases or events |
| min | minimum | phases |
| mean | mean | phases |
| std | Square root of the variance from the mean. Not yet tested. | phases |
| minframe | frame position, where the minimum is reached | phases |
| maxframe | frame position, where the maximum is reached in the case of application to a „phase“, „frame at“ in the case of an event | phases or events |
| frames | Count of total frames of a phase. These includes frames at which some values can be NaN. Not yet tested. | phase |
| dist |
Definition of Scopes
The scope defines event-types or phase-types, for which the parameters are determined. Have a look at the following table to learn about its attributes.
| Attribute name | Description |
|---|---|
| id | id to determine between different scopes, typically numbers starting with zero |
| name | name of phase- or event-type |
| type | „phase“ or „event“ |
Each scope can include one or more variables and properties:
<variable id="LeftScapulaPositionShiftLengthMax" name="LeftScapulaPositionShiftLengthMax"/> <property path="TRIAL_INFO/cycle_nr"/>
Definition of Processing Groups
This feature is used only, if datamining is based on a folder.
| Attribute name | Description |
|---|---|
| id | name of processing group as defined in the job config file |
| fileSuffix | If datamining is based on all trials of an selected folder, the trial data is read from the file with the given file suffix. |
| selected | true or false, depending of the trials defined by this group are included or excluded from the datamining. |
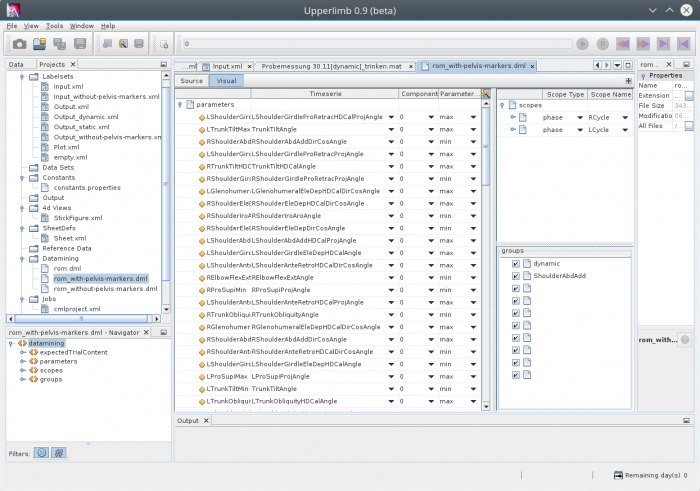
Visual view for the configuration
There is also a visual view available, to edit the configuration:
Dataminng based on such a configuration can be applied automatically on lists of datasets or on a selected dataset only.
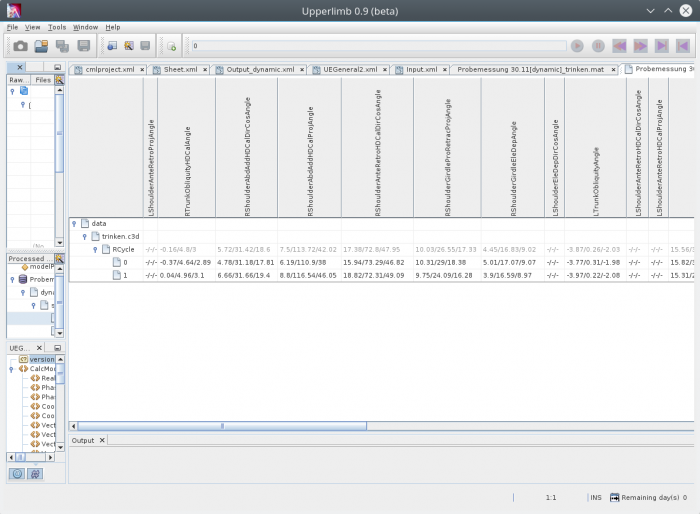
Visual view of the resulting data
Data can be selected by left-click on a trial child node of a processed d3d-file. Right-click
on the selected node opens the context-menu and you can execute the action „Open Datamining sheet“. This opens a View to min/rom/max of all timeseries which are defined as available in the configuration for all phases as defined in the configuration.
Export of CSV-files
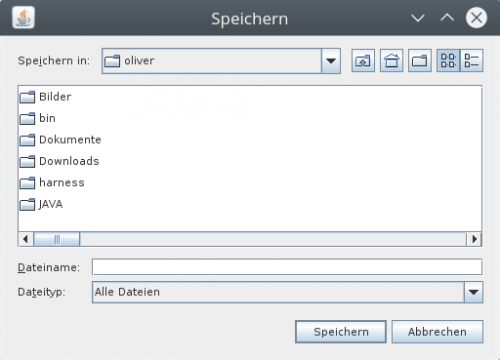
In the „visual view of resulting data“ you can right-click on the datas root node to open a context menu. The action „export“ opens a file chooser dialog.
You can choose the path and the file, where to export the data as a csv-file. The exported file can be opened with spreadsheet calculation software. In the case of Excel you can use the suffix „.xsl“ and it can be opened directly. Keep in mind that this is typcically a subset of the data shown in the view.
Known Issues
- Event- and Phase-Type-Names are shown as „scope type“ in the exported exelsheet and also as node name in the node-hierarchie of the UI. It is allowed to use the same name as event and phase type name. But in this case they can not be distinguished. As a workaround use only different names. In a future release the type „event“ or „phase“ should be added as a suffix between brackets, e.g. RGrasp[event] amd RGrasp[phase].